Abstract: While diagnostic problem solving is in principle
well understood, building and maintaining systems in large domains
cost effectively is an open issue. Various methods have different
advantages and disadvantages making their integration attractive.
When switching from one method to another in an integrated system,
as much knowledge as possible should be reused. We present an
inference structure for diagnostic problem solving optimized with
respect to knowledge reuse among methods motivated by experience
from large knowledge bases built in medical, technical and other
diagnostic domains.
1. Introduction
Diagnostic Problem Solving probably is the best understood subdomain of knowledge (expert) systems [Puppe 93, Breuker & Velde 94]. Diagnostic (classification) problems can be characterized as the selection of solutions from a preenumerated set of alternatives (diagnoses) for a problem described by observations (symptoms). Typical domain types (see [Puppe 93, chap. 13.2]) are medical diagnosis and therapy selection, object identification (e.g. plants, rocks), fault finding for technical devices and processes (e.g. car engines, printing machines), assessment (e.g. credit granting or legal advice) and precedence selection (e.g. looking for the best product). There are a variety of successful problem solving methods for categorical, heuristic, causal or case-oriented knowledge requiring different representations ([Puppe 96]; for variants not mentioned here, see e.g. [Benjamins 95]):
At least in principal, all methods can be applied to all classification
domains with one important exception: Functional and behavioral
classification are only applicable for fault finding (troubleshooting)
by searching for differences between the correct and a malfunction
of a system. Classification domain types without such a model
are e.g. object identification, assessment and precedence selection
(see above).
While the principal problems of diagnostic modeling are quite
well understood, and for most domains routine approaches can be
offered, the cost effectiveness of large knowledge bases (with
more than 1000 symptoms and diagnoses) is a major issue. For a
company, the crucial question is whether the expected benefits
from routine use exceeds the costs incurred from building and
maintaining the knowledge base. Since the knowledge acquisition
costs increase exponentially with the degree of completeness of
knowledge bases, we cannot make the assumption [Fensel &
Benjamins 96, van Harmeln & ten Teije 98], that knowledge
bases are complete - neither with respect to the coverage of all
possibly relevant observations nor with respect to the correctness
and completeness of the knowledge drawing conclusions from them.
In the following we concentrate on interactive systems with the
user entering at least some of the symptoms of the case (as opposed
to embedded systems). The goal of interactive systems is not primarily
to solve problems automatically but to increase user performance
(see e.g. the discussion in [Berner et al. 94]). Therefore, we
have to accommodate for vague, incomplete and incorrect data -
the latter due to difficulties of the user with symptom recognition.
In particular with respect to different compromises between knowledge
and data acquisition effort and accuracy of the resulting product,
the various methods have different advantages and disadvantages.
In the following, we discuss general knowledge acquisition strategies
with an emphasis on the costs for building and maintaining a knowledge
base.
Knowledge bases can be built by knowledge engineers supported
by domain experts, by the domain experts themselves or by machine
learning techniques. The latter is the least expensive alternative.
It usually requires a data base with many cases, where the observations
are stored in a standardized form together with the correct solutions.
The statistical, case-based and neural classification methods
belong to this category. In addition there are inductive learning
techniques like the ID3-family of decision tree learning algorithms
or the star method for learning rules (for a thorough evaluation
see [Michie et al. 94]). However, these algorithms currently cannot
solve the knowledge acquisition problem for several reasons. First,
a large effort is necessary to produce detailed, standardized
documentation forms for data gathering, which is still left to
the expert. Second, the raw data is often too low level for learning
algorithms, so that experts need to add preprocessing knowledge
(which we call data abstraction). Third, it is often quite useful
to formalize basic domain knowledge in advance. Fourth and most
important, the quality of the generated knowledge is often poor
compared to expert knowledge. One common reason is the lack of
sufficient cases for generating useful abstractions to classify
rare diagnoses. While we do not expect that learning from cases
can produce knowledge bases of similar quality as possible with
experts, it can be a useful technique to support manual knowledge
acquisition.
Since knowledge acquisition with two kinds of highly paid specialists
- knowledge engineers and domain experts - is often too expensive,
we concentrate on the alternative of domain experts building their
knowledge bases largely on their own. The first requirement is
a shell with a convenient, easy to learn and fast to use knowledge
acquisition interface. Visual programming languages [Burnett
& Ambler 94] offer these features and are particularly well
suited for diagnostic problem solving, because most knowledge
can be represented declaratively. To enter their knowledge the
experts have to fill in forms, tables, graphs or hierarchies [Bamberger
et al. 97].
2. Experience with building large knowledge bases by experts
In this section, we report experience from three projects, where experts built large knowledge bases in medical, technical and other domains with our diagnostic shell kit D3 [Puppe et al. 96]. Thereby we emphasize the aspects of knowledge restructuring. D3 offers most of the above mentioned methods including categorical, heuristic, statistic, set-covering, functional and case-based diagnostic problem solving methods, and a complete graphical knowledge acquisition interface [Gappa et al 93]. Knowledge in D3 can be used for various purposes:
Plant classification
The plant classification system was built by the biologist Roman
Ernst [Ernst 96]. It covers the 100 most common flowering plants
of the "Mainfranken" region of central Germany. Each
plant is characterized by visible features describing flowers,
leaves and trunk only, that is, no special instruments are necessary
to recognize the features (see Fig. 1). Since they are sometimes
ambiguous and difficult to recognize, the method must be able
to deal with many low-quality features and be robust enough to
tolerate some misrecognized features. The decision tree approach
used in many plant classification books does not share these assumptions,
because they sometimes rely on features detectable only by special
instruments and it usually cannot recover from misrecognized features.
Therefore decision trees were discarded.
The most obvious and time-effective approach uses set-covering
knowledge stating the features for each plant with typicality
(frequency) values. However, our evaluations proved this method
to be very sensitive to misrecognized features. To make the system
more robust, it was necessary to state not only the typical features,
but also to what degree the other features would be untypical
for each plant. This knowledge is straightforwardly expressed
with heuristic rules, where each feature transfers positive or
negative evidence for almost each plant. In the final knowledge
base there are more than 10000 one-symptom-one-diagnosis rules,
part of them shown in Fig. 2. Since they are entered in D3 through
tables like those in Fig. 2 (containing already almost 100 relations),
the effort for knowledge editing including fine tuning of the
evidence took only about one person month for the expert. Evaluations
demonstrated greater robustness than the set covering knowledge
base allowing even 1-3 misrecognized features from about 40 without
significant deterioration of performance. However, there were
still problems with some plants because of their varying phenotypes
depending on their environment. Since empirical knowledge was
not readily available for the variations, a case-based component
was added allowing the system to remember unusual cases (but they
must be classified the first time without the system). Therefore,
knowledge about a similarity measure was added and the cases already
gathered for evaluation of the other methods were used as the
basic case set. The results of this evaluation were encouraging
taking the restricted case set (150 cases for 100 different plants)
available into account, but showed a significant lower performance
than the other two methods.

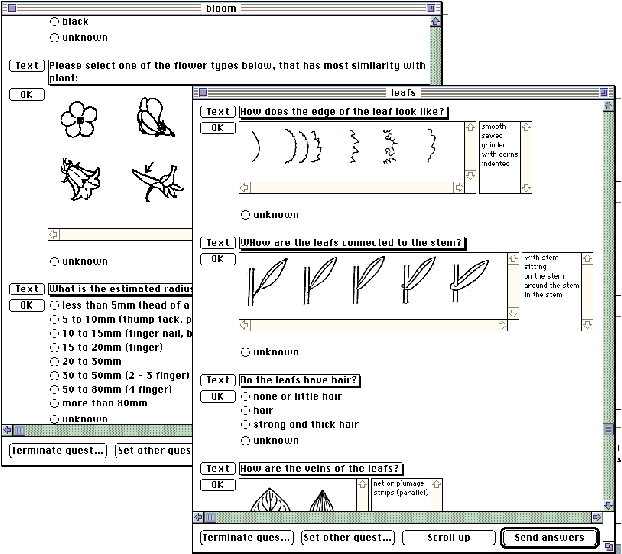
Fig. 1: User questionnaires for entering features of a plant.

Fig. 2. Input of simple heuristic rules with a table. Four
plants (column headers) are rated by the features in the lines.
In the marked box the following rule is entered: if the "perimeter
of the stock" (Stengelumfang) is "round and rather smooth"
(rund und eher glatt), there is sometimes (manchmal, p2) evidence
for "Taraxacum officinalis" (Wiesen-Löwenzahn).
Another focus of the work was to teach casual users to recognize
features correctly with the help of drawings and verbal descriptions.
Therefore, a training system was generated from the knowledge
base. It contained plant pictures presented to the user, who is
supposed to describe their features in terms of the terminology
of the system.
The biologist developed the knowledge base without knowledge engineering
support using the graphical knowledge acquisition facilities of
D3. A large amount of the total development time of six months
he spent with gathering, scanning and editing the plants and drawings
of typical features.
Service support for printing machines
The goal of this joint project with one of the world's leading
manufacturers of printing machines, Koenig&Bauer Albert AG
(KBA) in Wuerzburg, is the development of a diagnosis and repair
system both for the initial starting-up and subsequent service
support of printing machines [Landvogt et al. 97]. Since printing
machines are very large and complex, it was clear from the beginning,
that one huge knowledge base would be very difficult to maintain.
Therefore a concept was elaborated to split the knowledge base
in different modules (agents), which can be developed by different
experts more or less independently and which are able to cooperate
to solve cases (see chapter 7).
It quickly turned out, that a pure knowledge-based system was
not sufficient for this purpose, since company technicians expect
the computer to hold all relevant information including drawings,
parts lists, descriptions, videos and client contacts for the
particular machine. A large effort went in expanding the knowledge
representation of D3 to link formal and informal knowledge and
to integrate viewers for the different information sources.
With respect to continuous maintenance of the knowledge base,
it was a major goal of the project, that the KBA workers learnt
to develop the knowledge base on their own - ideally spending
one day per week for this task, while working the rest of the
week on their primary job. The organizational and qualificational
aspects of this "self-acquisition" (as opposed to knowledge
acquisition with knowledge engineers) are studied by a third project
partner IuK ("Institut für sozialwissenschaftliche Technik-Folgenforschung").
From a pure technical point of view, the self-acquisition did
work, for KBA experts indeed developed large knowledge bases on
their own with the graphical knowledge acquisition interface of
D3 (see table 1). However, for quality improvement general modeling
principles and technical tricks were discussed with "knowledge
consultants".
| Problem area | Symptom sets | Symptoms | Diagnoses | Rules |
| print unit JC | 46 | 365 | 258 | 1012 |
| print unit EC | 81 | 691 | 354 | 1877 |
| paper run | 6 | 80 | 103 | 495 |
| print quality | 10 | 38 | 69 | 164 |
| rell changer | 54 | 499 | 280 | 1608 |
| rabbet gadget | 7 | 198 | 101 | 697 |
| recontrol | 44 | 361 | 242 | 980 |
| sum | 248 | 2232 | 1407 | 6833 |
Table 1: Distributed knowledge
bases for printing machines: statistics in 1996.
Methodologically, the first approach was based on decision trees,
since that seemed most natural to the experts. The underlying
strategy is that they quickly generate suspects from the main
symptom and have specific questions and tests to find the diagnosis
(usually corresponding directly to a repair job). The decision
trees are rather broad and flat, with usually only 3 questions
in one branch. Accordingly, the key knowledge seemed to lie in
the sequence to gather symptoms. This knowledge was available
in a compiled form, so that it could be represented directly with
dialog-guiding rules. The main problem with this approach arose
from the technician's desire for early feedback after having entered
the first few symptoms. However, decision trees display results
only upon reaching a leaf in the tree. When there are many suspects,
often many branches must be explored until discovering the final
diagnosis. If the user is too impatient to finish a session, no
results at all might be displayed. The main requirement for the
system was to display all its suspects all over the time, i.e.
initially a large set of ranked hypotheses shrinking subsequently
with ruling-out of alternatives. The experts first tried to implement
this behavior with heuristic classification, because they preferred
the decision tree editor in D3 allowing also easy entering of
simple heuristic relations. To convince them to use the better
suited set covering classification required to extend that editor
to handle those relations as well. The basic knowledge structure
is quite simple: For every symptom, all diagnoses covering it
are stated. The set-covering problem solver also takes into account
confirmation or exclusion of a diagnosis by categorical knowledge.
At any time, it displays a list of diagnoses ranked to the degree
how well they explain currently known symptoms.
Rheumatology
The primary goals of the expert system RHEUMA from Prof. Schewe
(Munich university hospital) are to systematize collecting medical
history and clinical data, to assist non-specialist physicians
in rheumatic diagnostics, to document patient data, and to automatically
generate a patient report. RHEUMA comprises the approximately
70 most common diagnoses in rheumatology. The first approach was
based on heuristic classification, because it seemed to be most
effective in large medical domains. The results were quite encouraging:
In a prospective evaluation with 51 unselected outpatients coming
from a second clinic, it stated the final clinical diagnoses (which
was used as gold standard) in about 90% of the cases and focused
on the correct diagnoses in about 80% % (i.e. in about 90% of
the 90% cases with the final diagnoses) [Gappa et al 93]. Since
more than 1000 real cases were collected during development and
evaluation time, case-based knowledge about a similarity measure
was added to RHEUMA to exploit this data. However, due to multiple
diagnoses in many cases, the performance of the case-based classification
component scored significantly lower than heuristic problem solving.
In a recent extension the program is also used as a training system
with reversed roles of computer and user: The training system
presents case data step by step and invites the user to recognize
symptoms from pictures, order tests and suggest hypotheses. For
all these decisions, the training systems computes didactic feed-back
by comparing them to its own preference in the given diagnostic
context. The training system was evaluated in several mandatory
courses for students at Munich medical school with good results
[Schewe et al. 96]. While technically no modifications of the
diagnostic knowledge base were necessary, the heuristic rules
turned out to be too complicated for teaching purposes. To improve
its explanatory capabilities, a set-covering (pathophysiologic)
model is currently added to the heuristic and the case-based knowledge
base.
Both development and maintenance of the knowledge base was achieved
by the expert himself. He was even able to model his knowledge
according to different paradigms. Similar systems were developed
from other medical authors with D3 for large fields like neurology,
cardiology (an example of the definition of a rheuma-related differential
diagnosis from the cardiology knowledge base is shown in Fig.
3), hepatology, hematology as well as in many smaller domains
like thyroid diseases or acute abdominal pain [Puppe et al. 95].
| 1. Malar rash | |
| 2. Discoid rash | |
| 3. Photosensitivity | |
| 4. Oral ulcers | |
| 5. Arthritis | |
| 6. Serositis | |
| 7. Renal disease | a. > 0.5 g/dl proteinuria, or |
| b. >= 3+ dipstick proteinuria, or | |
| c. Cellular casts | |
| 8. Neurologic disease | a. Seizures, or |
| b. Psychosis (without other cause) | |
| 9. Hematologic disorders | a. Hemolytic anemia, or |
| b. Leukopenia (< 4000/µL), or | |
| c. Lymphopenia (< 1500/µL), or | |
| d. Thrombocytopenia (< 100.000 µL) | |
| 10. Immunologic abnormalities | a. Positive LE cell preparation, or |
| b. Antibody to native DNA, or | |
| c. Antibody to Sm, or | |
| d. False-positive serologic test for syphilis | |
| 11. Positive antinuclear antibody (ANA) |

Fig. 3. Definition of
"Systemic Lupus Erythematosus" (SLE), which requires
that at least 4 of the 11 criteria are met (upper part) and its
graphical knowledge acquisition (lower part) in two versions with
complex (A) and simple (B) n-of-m rules. The leftmost rule in
both versions represent the definition stated in the upper part
with a definite ("defi..." in the top most line) conclusion.
The other two rules of each part infer an evidence category (probable
or possible). The rules in (A) and (B) represent the same content,
but in (B), 4 supplementary rules are necessary for inferring
the symptom interpretations stated in the last 4 lines (D; renal
criterion, neurologic criterion, hematologic criterion and immunologic
criterion), which are omitted. Such a rule might be "if hematologic
evidence is Thromocytopenia or Lymphopenia or Leukopenia or Hemolytic
Anemia, then infer Hematologic Criterion = positive". This
rule represents the same knowledge as expressed in the 4 lines
about hematological evidence (C) marked with "+O3" within
the complex n-of-m rules (A).
3. Flexibility in knowledge acquisition
Successful self-acquisition of knowledge by domain experts in the above mentioned projects was made possible by D3's graphical knowledge acquisition component using graphical aggregation and abstraction techniques for managing the complexity of large knowledge bases [Gappa et al. 93, Bamberger et al. 97]. It not only simplifies the process of knowledge editing, but also makes the represented knowledge more transparent to the experts. But equally important is that the experts feel comfortable with the problem solving method used. Since there is often little time to train them, the starting point must be their experience with solving problems, i.e. there must be a match between the problem solving method of the computer and their own. Therefore a large variety of problem solving methods should be offered to them so they can pick up and configure the most suited. In the examples and in many other projects, switching or upgrading between different methods is necessary. While some methods are better suited for solving not so difficult problems, others require more initial modeling efforts but offer a higher degree of sophistication. Since the complexity of the problem is often not evident at the beginning, one wants to start with easy models and refine them if necessary. Therefore, from a representational point of view the knowledge for the different problem solving methods should be reused as much as possible to allow the domain experts to switch between methods with minimal effort. Some typical pathways through the problem solving methods are:
If switching to another problem solving method implied building
a completely new knowledge base, the costs would be in general
prohibitively high. However, there is much potential for reuse:
Taking into account this rather large potential of knowledge reuse,
we designed a combined inference structure for the different diagnostic
problem solving methods to exploit this potential. Besides allowing
a soft transition among methods, it also encourages building knowledge
bases with several methods in parallel to combine their advantages.
4. Knowledge sharing among diagnostic problem solving methods
The obvious solution for knowledge reuse is to modularize problem
solving methods according to reusable knowledge types. Hence we
introduced separate modules for data gathering, data abstraction,
categorical valuation of diagnoses and non-categorical valuation
of diagnoses. It is only the latter that differs among different
methods. Furthermore, it is desirable that all models share the
terminology. In addition, the diagnostic profile, i.e. a list
of relevant features for a diagnosis, and the abnormality of observations
are represented as separate data structures supporting knowledge
representations of different methods. In the following, we focus
on how different diagnostic might share and reuse that knowledge.
A standardized representation of observations and solutions might
already seem sufficient for case based classification methods
(in addition to cases). Additional useful knowledge includes the
similarity measure, the general information about abnormalities
and data abstractions. Diagnostic profiles might also be useful
for adding diagnosis-specific adaptations of the contribution
of certain observations to the overall similarity between cases.
Categorical valuations of diagnoses should take preference over
any similarities between cases. Otherwise, that knowledge would
have to be coded within the similarity measure, e.g. to prevent
a case of a male patient to be compared to a case of pregnancy.
Since case-based reasoning is not very well suited to support
data gathering (in particular when mostly unspecific symptoms
are known so far), it greatly gains by using compiled data gathering
knowledge.
Heuristic and set-covering knowledge can exploit nearly all reuse
facilities listed above: terminology, data abstraction, compiled
and hypothetico-deductive data gathering knowledge, categorical
knowledge for diagnostic reasoning, and diagnostic profiles. For
example, after implementation of this knowledge for heuristic
classification, only the particular formalization of the heuristic
rules based on the diagnostic profiles has to be reformulated
by set-covering relations (and vice versa). In addition, set-covering
classification needs knowledge about abnormalities of observations.
In troubleshooting domains, set covering knowledge can often be viewed as a compiled form of functional models:
- Discrepancies (functional) correspond to abnormal observations (set covering),
- abnormal component states (functional) correspond to primary diagnostic causes (set covering),
- abnormal parameters of materials (functional) correspond to
intermediate states (set covering).
To maximize knowledge reuse, we conceptualized and implemented
the functional model based on these correspondings as an extension
of the set-covering module [Kohlert 96].
Statistical classification mainly uses knowledge about a priori
probabilities and conditional probabilities of effects given the
causes. Both are computed from large case sets. They can reuse
knowledge about terminology, data abstraction, data gathering
and categorical knowledge for diagnostic reasoning. Given this
knowledge, only a subset of independent symptoms and mutually
exclusive diagnoses must be added for the straightforward application
of the Theorem of Bayes. Bayesian nets require in addition the
specification of the net topology, which is based on the same
principles as in set covering classification and therefore can
be reused from the diagnostic profiles.
For categorical classification, no special knowledge besides the
shared knowledge parts is necessary: Decision trees and decision
tables (by definition) contain only categorical knowledge for
diagnostic reasoning, and decision trees in addition use compiled
data gathering knowledge.
Fig. 4 summarizes an integrated inference structure for diagnostic
problem solving combining problem solving methods for categorical,
heuristic, statistic, set-covering, functional and case-based
classification optimized for knowledge reuse. Knowledge not specific
for a certain problem solving method is separated out as independent
modules (1.1 - 1.4 in fig. 4) being (re)used by any problem solving
method. Therefore, the specific part of a problem solving method
can be reduced to its "non-categorical valuation of diagnoses"
(1.5 in fig. 4). But even among these modules some reuse of knowledge
is possible as mentioned above (diagnostic profiles, abnormality
of observations), but not expressed in the inference structure.
A detailed description of the tasks and methods is given in the
appendix.

Fig. 4. Combined inference structure for categorical, heuristic, statistic, set-covering, functional and case-based classification in an and/or graph. Or-nodes, from which one or more successors can be selected, represent methods and are printed in ovals and italics. And-nodes denote tasks. The appendix presents details for the modules including informal description, input, output and necessary knowledge.
5. Comparison with other inference structures
Our modularization of diagnostic problem solving methods is distinguished
from others in its breadth of methods and in its emphasis of knowledge
reuse among the methods. In the following we compare it in some
detail with the inference structure presented in [Benjamins 95],
because this is based on a rather broad survey on the literature
of diagnostic systems. The three main tasks are symptom detection,
generation of hypotheses for the symptoms detected and finally
discrimination of the hypotheses. Symptom detection can be viewed
as a classification task in itself or may be done by the user
or by comparing expected with observed values. Expected values
can be looked up in a data base or predicted by a simulation.
For the comparison task, many methods exists ranging from an exact
comparison (if expected and observed value are identical, the
symptom is considered normal, otherwise abnormal) to various abstractions.
For the hypothesis generation task Benjamins offers only two
methods: compiled and model-based. Compiled methods consists of
the three tasks symptom abstraction, association of (abstracted)
symptoms with diagnoses and a probability filter. Model-based
methods consists of three methods too: Find possible causes (contributors)
for each abnormal symptom, combine the contributors to sets of
hypotheses being able to explain all abnormal symptoms and filter
out inconsistent hypotheses by simulating their effects in the
model. If there are several hypotheses left, additional observations
are gathered in order to discriminate between the hypotheses.
A summary of the inference structure is shown in fig. 5.
In contrast to the six problem solving methods in Fig. 4, Fig 5 contains only two: compiled and model-based, with many variants. The correspondences of the main steps are:
The main differences concern the emphasis of categorical knowledge.
While nearly lacking in the inference structure of [Benajmins
95], in our modularization it comes in many forms: standardized
test indication, plausibility check of answers, inferences of
follow-up questions, data abstraction, categorical valuation of
diagnoses, filter for symptoms and detection of abnormal symptoms.
If categorical knowledge exist, it should be applied with priority,
even for model-based problem solvers. This holds in particular,
if one cannot guarantee the completeness of the knowledge base
and therefore cannot apply the closed world assumption (e.g. reasoning
by elimination).
Another main difference is in the symptom detection task. Our
modularization lacks a step to generate expectations by prediction
(2.1.1.1.2), but instead offers the general mechanism of data
abstraction (1.3) and the detection of abnormal symptoms (1.5.5.1).
The latter is roughly equivalent to the look-up and compare step
(2.1.1.1.1 and 2.1.1.2). The data abstraction step is available
in [Benjamins 95] in the compiled hypothesis generation method
only. However, its mechanisms including arithmetic, aggregation
and abstraction are often powerful enough to detect abnormal symptoms
(e.g. inferring whether a measurement value like energy consumption
in a technical device is too high or computing the shock-index
from heart beat frequency and blood pressure). One might argue,
that a prediction model is needed anyway for model based diagnosis
in the hypothesis generation step, and therefore the respective
knowledge acquisition effort cannot be avoided. However, we found
that generating symptom expectations often requires a more detailed
model than generating and filtering hypotheses. Typical applications,
where it is not necessary to predict symptom expectations, comprise
not only medical diagnosis, but also technical diagnosis, if one
can built upon high level observations from e.g. automatically
generated error messages or humans. Of course, this does not hold
in all technical domains like e.g. circuit diagnosis, where prediction
is necessary for detection of abnormal symptoms. It should be
noted, that e.g. statistical and case-based classification are
difficult to apply in such domains.

Fig. 5. Condensed inference
structure for diagnosis from [Benjamins 95] compared with table
2 (in parentheses). "..." indicates further refinements
in [Benjamins 95].
A minor difference is the kind of modularization of the model based hypothesis generation method in both approaches. The three steps in "find contributors, transform to hypothesis set and prediction based filtered" (2.2.2.1 - 3) in [Benjamins 95] are distributed over the two steps "best cover based on single and multiple diagnoses" (1.5.4.3 - 4 and 1.5.5.3 - 4). The reason for our distinction is, that it is quite efficient to compute the cover for each relevant single hypothesis taking into account both covering and prediction, and the best single hypotheses can use the generalized knowledge for selecting the next test with the best costs/benefits ratio (1.1.3). The computational expensive step of computing multiple diagnoses for the best complete cover is often best done based on the results of the single cover step at the end or on explicit demand of the user.
6. Discussion
Different diagnostic methods offer different theoretical and practical advantages: they use different knowledge (categorical, empirical, case-based or causal) and they have different tradeoffs (e.g. knowledge acquisition time versus diagnostic accuracy, effort for building the initial prototype versus continuous maintenance) and explanation capabilities. Therefore, a major design goal for the diagnostic shell kit D3 - besides offering the most important methods - was to allow reuse of as much knowledge as possible when switching from one method to another. D3`s modularized inference structure is optimized towards this goal. All methods reuse knowledge about terminology and taxonomy of symptoms and diagnoses, local and global data gathering strategies (test selection and follow-up questions), plausibility checks, data abstraction and categorical valuation of diagnoses. Other knowledge structures shared by some methods are symptomatic profiles for diagnoses as well as knowledge about the importance and abnormality of symptom values. The knowledge representations and their visualization by graphical knowledge editors encouraged domain experts to represent their knowledge according to different paradigms. While this is useful for finding out the best method for both the domain and the mental model of the domain expert, it is even more useful for combining the advantages of different models. In medium or large medical domains such as rheumatology or cardiology this might be:
A further step in knowledge reuse is to take more advantage of
different problem solvers about the same domain, e.g. a heuristic,
set-covering and case-based reasoner. If they propose different
solutions for the same case, coming up with a combined solution
requires meta-knowledge about the general quality of the knowledge
bases and about the assessments of the individual solutions. Cooperation
can also take place during the problem solving process, e.g. using
heuristic or case-based knowledge for fast hypothesis generation
being tested by e.g. a simulation in a functional model.
A more ambitious step is the cooperation of independently developed knowledge bases like e.g. for different medical domains or different aggregates of large technical systems such as printing machines [Bamberger 97]. This distributed problem solving approach requires knowledge about:
Distributed problem solving is an important step to reduce the
size and thereby the complexity of knowledge bases and to increase
reuse, because for example a knowledge base for X-ray interpretation
can be consulted by many different medical knowledge bases.
7. References
Bamberger, S. (1997). Cooperating Diagnostic Expert Systems to Solve Complex Diagnosis Tasks, in Proc. KI-97, Brewka, G., Habel, C. and Nebel, B.: Advances in Artificial Intelligence 1997, LNAI 1303, Springer, 325-336.
Bamberger, S., Gappa, U., Kluegl, F. and Puppe, F. (1997). Komplexitätsreduktion durch grafische Wissensabstraktionen [Complexity reduction by graphical knowledge abstractions], in Mertens, P. and Voss, H. (eds.): Proc. of XPS-97, 61-78, infix.
Benjamins, R. (1995). Problem-Solving Methods for Diagnosis and their Role in Knowledge Acquisition, International Journal of Expert Systems: Research and Application.
Berner, E. et al. (1994). Performance of four Computer Based Diagnostic Systems, New England Journal of Medicine 333, 1792-1796.
Breuker, J and van de Velde, W. (eds. 1994). COMMONKADS Library for Expertise Modeling - Reusable Problem Solving Components, IOS-Press.
Burnett, M and Ambler, A. (1994). Declarative Visual Programming Languages, Journal of Visual Languages and Computing 5, Guest Editorial, 1-3.
Ernst, R. (1996). Untersuchung verschiedener Problemlösungsmethoden in einem Experten- und Tutorsystem zur makroskopischen Bestimmung krautiger Blütenpflanzen [Analysis of various problem solving methods with an expert and tutoring system for the macroscopic classification of flowers], Master thesis, Wuerzburg university, Biology department.
Fensel, D. and Benjamins, R. (1997). Assumptions in Model-Based Diagnosis, AIFB-Internal Report 360, Karlsruhe University.
Gappa, U., Puppe, F., and Schewe, S. (1993). Graphical Knowledge Acquisition for Medical Diagnostic Expert Systems, Artificial Intelligence in Medicine 5, 185-211.
Gappa, U. and Puppe, F. (1998). A Study of Knowledge Acquisition - Experiences from the Sisyphus III experiment of rock classification, To appear in Proc. of 12th Banff Knowledge Acquisition for Knowledge-Based Systems Workshop (KAW-98).
Kohlert, S. (1995). Eine Shell für minimale funktionale Klassifikation am Beispiel der Druckmaschinendiagnostik [a shell for minimal functional classification exemplified on printing machine diagnosis], Master Thesis, Wuerzburg University, Informatics department.
Landvogt, S., Bamberger, S., Hupp, J., Kestler, S. and Hessler, S. (1997). KBA-D3: ein Experten- und Informationssystem zur Fehlerdiagnose von Druckmaschinen, [KBA-D3: an expert and information system for troubleshooting in printing machines] in Mertens, P. and Voss, H. (eds.): Proc. of XPS-97, 167-170, infix.
Michie, D., Spiegelhalter, D., and Taylor, C. (eds., 1994). Machine Learning, Neural and Statistical Classification, Ellis Horwood.
Puppe, B., Ohmann, C., Goos, K., Puppe, F. and Mootz, O. (1995). Evaluating Four Diagnostic Methods with Acute Abdominal Pain Cases, Methods of Information in Medicine 34, 361-368,
Puppe, F. (1993). Systematic Introduction to Expert Systems - Knowledge Representations and Problem Solving Methods, Springer.
Puppe, F., Gappa, U., Poeck, K. und Bamberger, S. (1996). Wissensbasierte Diagnose- und Informationssysteme [Knowledge based diagnosis and information systems], Springer.
Schewe, S., Quack, T., Reinhardt, B., and Puppe, F.(1996). Evaluation of a Knowledge-Based Tutorial Program in Rheumatology - a Part of a Mandatory Course in Internal Medicine, in: Proc. of Intelligent Tutoring Systems (ITS-96), Springer, 531-539.
Van Harmelen, F. and ten Teije, A.(1998). Characterizing Problem
Solving Methods by Gradual Requirements: Overcoming the yes/no
Distinction, in Proc. KEML-98, 8th Workshop on Knowledge
Engineering: Methods & Languages, 1-15, Univ. Karlsruhe, 1998.
(URL: http://www.aifb.uni-karlsruhe.de/WBS/dfe/keml98).
Appendix: Description of Tasks and Methods for Fig. 4
| 1. Classification (Diagnosis) |
| Description: Selection of one or more diagnoses based on symptoms which may be given or which must be requested. |
| Input: Observed Symptoms. |
| Output: Diagnoses. |
| Knowledge: s. subtasks (1.1 - 1.5). |
| 1.1 Test selection |
| Description: Selection of tests, which should be performed for clarification of the case. |
| Input: Currently known symptoms and suspected and confirmed diagnoses. |
| Output: Tests (question sets). |
| Knowledge: s. subtasks (1.1.2 + 1.1.3). |
| 1.1.2 Standardized indication |
| Description: Selection of tests based on standardized procedures. |
| Input: Facts. |
| Output: Tests. |
| Knowledge: 1. Rules for indication of tests from constellation of facts. |
| 2. Rules for contraindication of tests from constellation of facts. |
| 3. Cost categories for tests. If more than on tests is indicated, the test with the lowest cost category will be selected. |
| 1.1.3 Costs/benefits analysis |
| Description: Selection of tests for best clarification of suspected diagnoses with the lowest costs. |
| Input: Suspected diagnoses (+ facts) |
| Output: Test |
| Knowledge: It is specified, which diagnoses how well will be clarified by a test. In addition the static (represented by an attribute) and dynamic (represented by rules) costs of a tests are stated. |
| 1.2 Data gathering |
| Description: Data gathering from the user or from external data sources. |
| Input: Test (question set). |
| Output: Assignment of values to the questions of the test. |
| Knowledge: s. subtasks (1.2.1 - 1.2.3). |
| 1.2.1 Interviewer component |
| Description: User interface for entering data or program interface for external data sources, which is not treated in this paper. |
| 1.2.2 Plausibility check of answers |
| Description: Plausibility check of input data. |
| Input: Answers to questions |
| Output: o.k. or error message. |
| Knowledge: 1. valid value ranges for each question |
| 2. Constraints for recognition of inconsistencies between different answers. |
| 1.2.3 Inference of follow-up questions |
| Description: Determination, whether follow-up questions should be asked. |
| Input: Answers to questions |
| Output: New questions |
| Knowledge: Rules for inferring questions. |
| 1.3 Data abstraction |
| Description: Derivation of symptom abstractions. They include arithmetic formulas for combining quantitative data, the abstraction of quantitative to qualitative data, and the aggregation of qualitative data. |
| Input: Symptoms |
| Output: values for symptom abstractions. |
| Knowledge: 1. Terms for symptom abstractions and their ranges. |
| 2. Rules for inferring the symptom abstractions, whose action part may be a number, a formula or a qualitative value. |
| 3. Schemes for transformation of numerical to qualitative values. |
| 1.4 Categorical valuation of diagnoses |
| Description: Confirmation or exclusion of diagnoses with categorical rules. |
| Input: Facts |
| Output: Confirmed or excluded diagnoses. |
| Knowledge: Rules, whose action part establish or exclude a diagnosis . |
| 1.5 Non-categorical valuation of diagnoses |
| Description: Valuation of diagnoses based on symptoms with vague knowledge. |
| Input: Facts |
| Output: Suspected and/or confirmed diagnoses. |
| Knowledge: Depends on problem solving methods (s. 1.5.1 - 1.5.5) |
| 1.5.1 Heuristic valuation |
| Description: Diagnoses are confirmed, if their accumulated evidence surpasses a threshold or if they are better than their competitors (differential diagnoses). |
| Input: Facts |
| Output: Suspected and/or confirmed diagnoses. |
| Knowledge: 1. Rules, whose action part contains a diagnoses and a positive or negative evidence category. |
| 2. A-priori frequency: Category from extremely rare via average to extremely common. |
| 3. Rules for dynamic correction of the a-priori frequency based on facts. |
| 4. Knowledge about direct competitors belonging to a diagnostic class, i.e. if the diagnostic class is confirmed, the highest rated competitor inside the class should be confirmed. |
| 1.5.1.1 Inference of suspected diagnoses |
| Description: A diagnosis is suspected, if its accumulated evidence is higher than a threshold and lower than another threshold. Suspected diagnoses are used as input for the cost benefits analysis. |
| 1.5.1.2 Inference of confirmed diagnoses |
| Description: A diagnosis is confirmed, if its accumulated evidence is more than the second threshold for suspected diagnoses or if it is the leader of a competitor class. Confirmed diagnoses can be treated in rule preconditions like symptoms. |
| 1.5.2 Simple statistic valuation |
| Description: Determination of the best diagnosis from a complete and exclusive set of alternatives with the Theorem of Bayes. |
| Input: Symptoms (which should be independent of each other). |
| Output: Diagnosis. |
| Knowledge: A-priori probability of diagnoses P (D) and conditional probabilities P (D/S) for all singular symptom/diagnosis combination, which should be extracted from large representative case sets. |
| 1.5.3 Case based valuation |
| Description: For a new case without solution one or more most similar cases from case set are extracted. If the similarity is high enough, the diagnoses from the old case(s) are transferred to the new case. The similarity of two cases is computed from the similarity of corresponding symptoms. |
| Input: Symptoms and (optional) a view, i.e. an abstraction layer. |
| Output: Most similar case(s) and their diagnoses. |
| Knowledge: For each symptom: Abstraction layer, weight, abnormality of its values, similarity type, similarity information. For each diagnosis it is possible, to state a specific weight and similarity information per symptom overriding the defaults. |
| 1.5.3.1 Filter for symptoms |
| Description: Some symptoms are filtered out, which should not be used for computation of similarity. |
| Input: Symptoms and view (abstraction layer). |
| Output: Subset of symptoms. |
| Knowledge: Abstraction layer of symptoms, which is compared with the input abstraction layer. |
| 1.5.3.2 Preselection of cases |
| Description: All cases are preselected, which are at least in one important symptom similar to the new case. |
| Input: Symptoms |
| Output: Cases |
| Knowledge: Important symptoms are determined by their weight and their abnormality. |
| 1.5.3.3 Elimination of cases with impossible diagnoses |
| Description: From the preselected cases those are sorted out, which have diagnoses being incompatible with the symptoms of the new case based on the categorical valuation. |
| Input: Cases, excluded diagnoses. |
| Output: Subset of cases. |
| Knowledge: Categorical valuation (1.4). |
| 1.5.3.4 Complete comparison of cases |
| Description: Determination of similarity between two cases as percentage number. |
| Input: New case and n old cases for comparison. |
| Output: For each old case the percentage of similarity. |
| Knowledge: s. 1.5.3. |
| 1.5.3.5 Determination of diagnoses |
| Description: From a list of similar cases, the solution for the new case is determined. There are several options: Take over of the diagnoses from the most similar case, if distance is high enough; majority vote of several very similar cases, if some of them have the same diagnoses; or more complicated transfers. |
| Input: Ordered list of cases with similarity rating. |
| Output: Diagnoses. |
| Knowledge: Diagnoses of the old cases and similarity ratings. |
| 1.5.4 Set covering valuation |
| Description: Selected is the diagnosis or set of diagnoses, which can cover (predict, explain) as much as possible of the facts and predict as less as possible unobserved facts taking into account the general frequency of the diagnoses. Completeness of the covering relations is not assumed. |
| Input: Symptoms (and optional abstraction layer). |
| Output: Diagnosis or set of diagnoses. |
| Knowledge: 1. Rules, whose precondition are a diagnosis and whose action are facts being covered by that diagnosis. The rules may have assumptions and frequency information. |
| 2. Abstraction layer, weight and abnormality of symptoms. |
| 3. A-priori frequency: Category from extremely rare via average to extremely common. |
| 4. Rules for dynamic correction of the a-priori frequency based on facts. |
| 5. Weight and abnormality information for symptoms. |
| 1.5.4.1 Detection of abnormal symptoms |
| Description: Symptoms are rated according their abnormality. |
| Input: Symptoms. |
| Output: Abnormality rating of symptoms. |
| Knowledge: Knowledge about the abnormality of the symptom values. |
| 1.5.4.2 Elimination of impossible diagnoses |
| Description: All diagnoses excluded by categorical reasoning are eliminated. |
| Input: Diagnoses. |
| Output: Subset of diagnoses. |
| Knowledge: Knowledge of categorical valuation (1.4). |
| 1.5.4.3 Best cover based on single diagnoses |
| Description: Each diagnosis checks its rating how well is covers the symptoms. The diagnosis are sorted and the best diagnoses are treated like suspected diagnoses for test selection (s. 1.1.3). |
| Input: Symptoms. |
| Output: Ordered list of diagnoses. |
| Knowledge: s. 1.5.4. |
| 1.5.4.4 Best cover based on multiple diagnoses |
| Description: Computation of a set of diagnoses, which together explain the symptoms better than a single diagnosis and which are according to their combined frequencies not too improbable. |
| Input: Symptoms and list of single diagnoses with their covering relations. |
| Output: Multiple diagnoses. |
| Knowledge: Frequency and covering relations of single diagnoses. |
| 1.5.5 Functional valuation |
| Description: Diagnoses are viewed as states of components and symptoms correspond to materials or signals between the components. The goal is to find those component states, which can predict (explain, cover) the normal and abnormal measurements of materials or signals. |
| Input: Measurement of materials and signals. |
| Output: Faulty component states. |
| Knowledge: Model of the normal behavior of a (technical) system, how input materials are transformed by components to output materials. The components have a normal state and various faulty states (diagnoses), whose behavior is described by rules, inferring values for the output materials from the input materials. The materials consist of parameters, which are represented as symptoms, if they can be measured, or by diagnoses, if they are used for further inferences. With these assumptions and transformations, the functional valuation can be viewed as an extension of the set covering valuation. |